News and Blogs
Comparison of Different DNA Testing Techniques and Their Applications

DNA testing techniques have revolutionized various fields, from criminal investigations to medical diagnoses, by providing valuable information about a person's genetic makeup.
In the UK, a recent court case was affected by an error at a DNA testing laboratory, Synlab Laboratory Services, which led to concerns about the reliability of DNA testing methods. The error resulted in around 1,800 drivers facing drug-driving convictions potentially having their cases quashed due to issues with samples analysed by Synlab between April 2019 and December 2020. The incident highlights the crucial role of accurate DNA testing techniques in legal proceedings and underscores the need to compare and evaluate different DNA testing methods and their applications carefully.
DNA testing techniques have become a crucial tool in criminal investigations, paternity tests, and identifying victims of disasters or crimes. However, different DNA testing methods have varying degrees of accuracy, sensitivity, and specificity, making it essential to understand their advantages and limitations. In this context, this article aims to provide a detailed comparison of different DNA testing techniques and their applications to help readers make informed decisions about which method to use for specific scenarios.
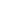
I. PCR-based DNA detection technique
PCR-based DNA detection technique is a molecular biology method used to amplify and detect specific DNA sequences. The Polymerase Chain Reaction (PCR) is a laboratory process that is widely used to create millions of copies of a specific DNA segment, allowing for easy detection and analysis. This technology has revolutionized DNA analysis and detection, as it can detect even small amounts of DNA in various samples, such as blood, saliva, tissues, and other body fluids.
The PCR-based DNA detection technique involves temperature-dependent enzymatic reactions that use DNA polymerase, primers, and deoxyribonucleotides (dNTPs) to amplify a specific DNA sequence. The process begins with denaturation, where the DNA double helix is heated to separate it into two single strands. Short DNA primers then bind to the complementary DNA strands flanking the target sequence. DNA polymerase then adds nucleotides to extend the primers, creating a new DNA strand that matches the template. This process is repeated multiple times through a series of temperature cycles, resulting in millions of copies of the target DNA sequence.
PCR-based DNA detection has numerous applications in various fields, such as medical diagnostics, genetic research, forensic science, and environmental monitoring. For instance, PCR-based DNA detection is widely used in medical diagnostics to identify infectious agents like viruses and bacteria, and to diagnose genetic disorders. In forensic science, PCR-based DNA detection is used to identify suspects or victims of crimes by analyzing DNA samples from crime scenes, including blood, semen, saliva, and hair. Additionally, this technique is used in genetic research to study gene expression, mutations, and genetic variations.
However, the PCR-based DNA detection technique has some limitations. One of the major limitations is its susceptibility to contamination, which can produce false positive results. Another limitation is the inability to amplify large DNA sequences or whole genomes.
The article "Risk assessment of false-positive quantitative real-time PCR results in food, due to detection of DNA originating from dead cells" by Petra Wolffs et al. discusses the risks of false-positive PCR results due to detection of DNA originating from dead cells. The article highlights that one of the main challenges concerning correct pathogen diagnostics by real-time PCR lies in the inability to distinguish between signals originating from viable cells and DNA released from dead cells. As a result, the risk of false-positive PCR results due to detection of dead cells is a major concern.
The study aimed to evaluate the effect of complex food samples on the persistence of chromosomal and plasmid DNA, using Yersinia enterocolitica as a model system. The study showed that in pork samples, there is a risk of false-positive PCR results due to detection of dead cells. This was confirmed in a quantitative study on cell death and signal persistence over a period of 28 days, employing three different methods, i.e. viable counts, direct qPCR, and finally, floatation, a recently developed discontinuous density centrifugation method, followed by qPCR. The study concluded that by using floatation as a sample treatment step prior to qPCR, the risk of false-positive PCR results due to detection of dead cells can be minimized.
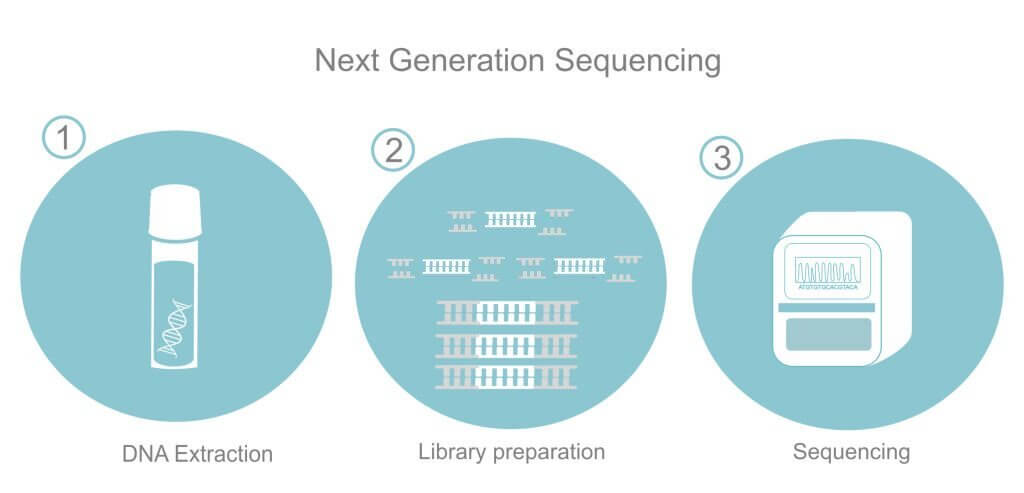
II. NGS-based DNA detection technique
NGS-based DNA detection technique is a next-generation sequencing method that allows for the analysis of multiple DNA sequences in a single run. This technology has revolutionized DNA analysis and detection, as it allows for high-throughput sequencing of DNA samples, making it possible to analyze large amounts of genetic data in a short amount of time.
NGS-based DNA detection involves a series of steps, including DNA extraction, library preparation, sequencing, and data analysis. DNA extraction involves the isolation of DNA from the sample, while library preparation involves the fragmentation of DNA and the addition of adapters to facilitate sequencing. Sequencing involves the use of NGS instruments, such as Illumina or Oxford Nanopore, to generate millions of sequence reads, which are then analyzed using bioinformatics tools.
NGS-based DNA detection has numerous applications in various fields, such as medical diagnostics, genetic research, and environmental monitoring. In medical diagnostics, NGS-based DNA detection is used to identify genetic mutations associated with diseases, such as cancer, and to develop personalized treatment plans. In genetic research, NGS-based DNA detection is used to study the genetics of diseases, population genetics, and evolution. Additionally, NGS-based DNA detection is used in environmental monitoring to detect microbial communities, identify plant and animal species, and monitor water quality.
However, NGS-based DNA detection has some limitations. One of the major limitations is the cost and complexity of the technology, which makes it less accessible than other DNA detection methods. Additionally, NGS-based DNA detection requires specialized equipment and expertise, which may limit its use in some settings.
The article "Costs of Next-Generation Sequencing Assays in Non-Small Cell Lung Cancer: A Micro-Costing Study" provides specific cost estimates for four next-generation sequencing (NGS) assays used for non-small lung cancer (NSCLC) diagnostics in Canada. Based on a case throughput of 500 cases per year, the per-sample cost for TruSight Tumor 170 Kit, QIASeq Targeted DNA Custom Panel and QIASeq Targeted RNAscan Custom Panel, Oncomine Focus, and HyperPlus/SeqCap EZ were CAD 1778, CAD 599, CAD 1100 and CAD 1270, respectively. The study also identifies the main cost drivers for these assays, with library preparation and sequencing accounting for 34-60% and 31-51% of costs, respectively. Additionally, the article cites previous cost studies, such as one that estimated the cost of targeted genomic sequence analysis of DNA from solid tumor specimens to range from USD 577.99 to USD 907.82 (CAD 816-1281) and another that estimated the per-sample costs for small and medium targeted gene panels to range from EUR 606-EUR 956 (CAD 1441-2273) and EUR 1137-EUR 3009 (CAD 2703-7154), respectively.
III. Microarray-based DNA detection technique
Microarray-based DNA detection technique is a powerful tool used to detect and analyze specific DNA sequences simultaneously. The method uses microarrays, also known as DNA chips or gene chips, which are glass or silicon chips with thousands or millions of probes that can bind to complementary DNA sequences.
Microarray-based DNA detection involves the hybridization of labeled target DNA with complementary probes immobilized on the microarray surface. The labeled target DNA is hybridized to the probes on the microarray, and the resulting fluorescence intensity is detected and analyzed using specialized software.
Microarray-based DNA detection has numerous applications in various fields, including medical diagnostics, genetic research, and forensic science. In medical diagnostics, microarray-based DNA detection is used to identify genetic mutations associated with diseases, such as cancer, and to develop personalized treatment plans. In genetic research, microarray-based DNA detection is used to study gene expression, gene regulation, and genetic variations. Additionally, in forensic science, microarray-based DNA detection is used to identify suspects or victims of crimes by analyzing DNA samples from crime scenes.
One of the advantages of microarray-based DNA detection is its ability to analyze multiple DNA sequences simultaneously, making it a high-throughput technique. This feature is particularly useful in applications where large numbers of samples need to be analyzed in a short amount of time. Additionally, microarray-based DNA detection is highly sensitive and specific, allowing for the detection of even small amounts of DNA in various samples, including blood, saliva, tissues, and other body fluids.
Microarrays have several significant drawbacks, including their high cost for a single experiment and the large number of probe designs based on sequences with low specificity. Furthermore, most commonly used microarray platforms only use one set of probes designed by the manufacturer, limiting control over the pool of analyzed transcripts. Additionally, microarrays have relatively low accuracy, precision, and specificity, and the experimental setup is highly sensitive to variations in hybridization temperature, genetic material purity and degradation rate, and the amplification process. These factors, among others, may have an impact on estimates of gene expression.

IV. Comparison of 3 DNA testing techniques
DNA detection technologies are critical tools in various fields, including medical diagnosis, genetic research, and forensic science. Different DNA detection technologies have their advantages and disadvantages, and the choice of technology depends on the application's needs. Here is a comparison of different DNA detection technologies in terms of their sensitivity, specificity, and cost, along with a specific example for each technique.
PCR-based DNA detection technique:
Pros:
High sensitivity: PCR technology can amplify the target DNA sequence and can detect very low amounts of DNA molecules.
High specificity: PCR technology has high specificity, and it only amplifies specific DNA sequences, minimizing the risk of false positives.
Low to medium cost: PCR-based DNA detection technology is relatively affordable compared to other DNA detection technologies.
Cons:
Limited scope: PCR can only amplify specific DNA sequences, which means it cannot detect unknown DNA sequences.
Vulnerable to contamination: The PCR process is susceptible to contamination, which can lead to false positives and inaccurate results.
Limited throughput: PCR can analyze a limited number of samples at a time, which can limit its usefulness for high-throughput applications.
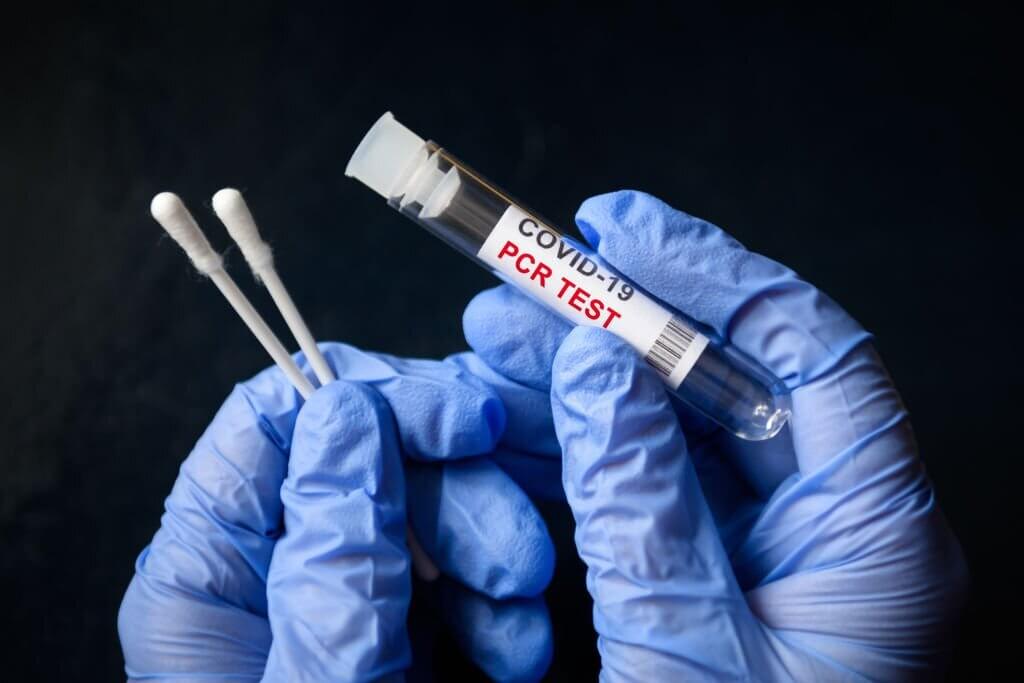
Example: PCR-based DNA detection technology is widely used in diagnosing infectious diseases, such as COVID-19. Polymerase chain reaction (PCR) tests are the most common type of diagnostic test used to detect the virus's genetic material in patient samples.
NGS-based DNA detection technique:
Pros:
High sensitivity: NGS technology can detect very low amounts of DNA molecules with high sensitivity.
High specificity: NGS technology has high specificity and can sequence the entire genome, making it an ideal technology for comprehensive genetic analysis.
High throughput: NGS can process a large number of samples simultaneously.
Cons:
High cost: NGS technology is expensive, and the initial setup cost can be high, including the specialized equipment and professional knowledge required.
Data analysis complexity: NGS produces large amounts of data that require extensive analysis, which can be challenging and time-consuming.
Errors: NGS can produce errors due to its complex nature and reliance on algorithms, which can impact accuracy.
Example: NGS-based DNA detection technology is used for cancer diagnosis and treatment. For example, in precision oncology, NGS technology is used to analyze cancer genomes, which can help identify genetic alterations that can be targeted by specific treatments.
Microarray-based DNA detection technique:
Pros:
High sensitivity: Microarray technology can detect even a single nucleotide variation with high sensitivity.
High specificity: Microarray technology has high specificity, and it can analyze multiple DNA sequences simultaneously.
High throughput: Microarray technology can process a large number of samples simultaneously.
Cons:
High cost: Microarray technology can be expensive, and the cost may vary depending on the number of probes used.
Limited scope: Microarrays can only analyze known DNA sequences, which can limit their use for discovering new genetic variations.
Data interpretation: Microarray data can be complex and require specialized knowledge for proper interpretation.
Example: Microarray-based DNA detection technology is used in genetic testing for diseases like cystic fibrosis. Microarrays can detect known genetic mutations associated with the disease, making them a useful tool for genetic testing and diagnosis.
In a word, different DNA detection technologies have their unique strengths and weaknesses, and choosing the appropriate technology depends on the application's needs. PCR is useful for applications that require high sensitivity and specificity at a relatively low cost, while NGS is ideal for comprehensive genetic analysis but can be expensive. Microarray technology is suitable for high-throughput analysis of multiple DNA sequences simultaneously.

V. The importance of proper sample collection and handling
Proper sample collection and preservation are crucial for accurate DNA test results. The quality of the sample collected can affect the sensitivity and specificity of the test, leading to incorrect or inconclusive results. Therefore, it is important to use appropriate sample collection methods and preserve the sample correctly.
Common sample collection methods include blood draws, oral swabs, and saliva samples. Blood draws are invasive and require a trained healthcare professional, but they provide high-quality DNA samples. Oral swabs and saliva samples are non-invasive and can be self-collected, making them more convenient. However, they may contain inhibitors that affect DNA quality, leading to lower sensitivity and specificity.
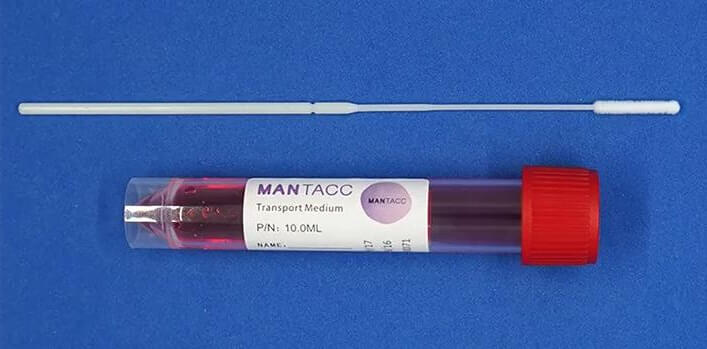
Mantacc DSK-F10-96A Disposable Sampling Kit is a convenient and reliable solution for nasal sample collection. The kit includes one nasal flocked swab and one non-inactivated transport media tube with buffer, designed to preserve the DNA sample during transport. The nasal flocked swab has an ergonomic and anatomic design that improves patient comfort and efficiency in cell specimen collection. The transport medium is formulated with antibiotics to inhibit bacterial and fungal growth, without affecting viruses, and can maintain organism viability for up to 48 hours at room or refrigerated temperature.
The DSK-F10-96A kit offers several advantages, including ease of use, safe sample transport, and compatibility with various diagnostic tests. The flocked swab's open fiber structure allows for superior sample elution, improving assay sensitivity, and increasing the assay's accuracy. The kit is also free of any PCR inhibitors, ensuring its compatibility with various diagnostic tests.
Studies have shown that the DSK-F10-96A Disposable Sampling Kit offers high accuracy, with a sensitivity of over 90%. Customers have reported high satisfaction rates, citing the kit's ease of use and reliability.
VI. Future outlook of DNA testing
DNA testing is a powerful tool that can reveal a lot of information about ourselves and our ancestors, as well as help solve crimes and identify victims. However, DNA testing also faces many challenges and opportunities in the future, as new technologies and applications emerge. Here are some of the main aspects of the future of DNA testing:
Emerging technologies: DNA testing techniques and technologies are constantly evolving and improving, making them faster, more sensitive, more accurate, and more affordable. Some of the emerging technologies include nanopore sequencing, which can sequence long DNA molecules in real time12; CRISPR-based DNA detection, which can use gene-editing enzymes to identify specific DNA sequences34; and DNA origami, which can fold DNA into complex shapes and structures for various purposes .
Applications: DNA testing has many applications in various fields, such as medicine, forensics, genealogy, and biotechnology. Some of the applications include personalized medicine, which can tailor treatments and drugs based on a person’s genetic profile ; forensic DNA phenotyping, which can predict a person’s physical appearance and ancestry from DNA evidence ; genetic genealogy, which can trace a person’s family history and ethnicity from DNA samples ; and synthetic biology, which can create new biological systems and organisms from DNA .
Challenges and opportunities: DNA testing also poses many challenges and opportunities for the society, the law, and the ethics. Some of the challenges include privacy and security, which can protect a person’s genetic information from unauthorized access and misuse ; quality and standards, which can ensure the reliability and validity of DNA testing results and methods ; and education and awareness, which can inform the public and the professionals about the benefits and limitations of DNA testing . Some of the opportunities include justice and human rights, which can use DNA testing to exonerate the innocent and identify the perpetrators of crimes and human rights violations ; health and wellness, which can use DNA testing to prevent and treat diseases and improve the quality of life ; and innovation and discovery, which can use DNA testing to explore new frontiers of science and technology.

VII. Conclusion
Different DNA detection technologies have varying strengths and weaknesses in terms of sensitivity, specificity, accuracy, cost, and throughput. We can examine how shared DNA and family connections play a crucial role in the early detection and prevention of diseases like breast cancer. By analyzing the genetic similarities within families, researchers can identify specific gene mutations and patterns that contribute to an increased risk of breast cancer. This knowledge enables healthcare professionals to provide personalized care and monitor those who are more susceptible to the disease, ultimately improving patient outcomes and overall public health. Proper sample collection and preservation are crucial for accurate DNA test results, and choosing the appropriate DNA detection technology depends on the specific application and desired level of sensitivity and specificity. The recent error at Synlab Laboratory Services highlights the importance of reliable DNA testing and the need for due diligence to ensure evidence entering the courts is reliable. The future of DNA testing looks promising, with the emergence of new technologies and applications that have the potential to revolutionize various fields, but their development and application come with both challenges and opportunities, and the accuracy and reliability of DNA testing results depend on proper sample collection, analysis, and interpretation, as well as appropriate regulation and ethical considerations.
Want to learn more about our forensic sampling kits?
Mantacc provides an extensive selection of top-quality forensic sampling products, including nasopharyngeal swab, oropharyngeal swab, and applicators. Our swabs are designed to ensure maximum patient comfort and are available in various materials and sizes. If you can't find what you're looking for, we also offer a professional customization service. Check out our collection by clicking on the link provided or contact our sales team by clicking on the image below.
Find products: forensic sampling kit, Mantacc Test Kits
Reference
1.Petra Wolffs, Börje Norling, Peter Rådström, Risk assessment of false-positive quantitative real-time PCR results in food, due to detection of DNA originating from dead cells, Journal of Microbiological Methods,
Volume 60, Issue 3, 2005, Pages 315-323, ISSN 0167-7012, https://doi.org/10.1016/j.mimet.2004.10.003.
2.Kumar S, Bennett A, Campbell PA, Palidwor G, Lo B, Perkins TJ, Nochaiwong S, Sekhon HS, Stewart DJ, Thavorn K. Costs of Next-Generation Sequencing Assays in Non-Small Cell Lung Cancer: A Micro-Costing Study. Curr Oncol. 2022 Jul 23;29(8):5238-5246. doi: 10.3390/curroncol29080416. PMID: 35892985; PMCID: PMC9330154.
3.Bumgarner R. Overview of DNA microarrays: types, applications, and their future. Curr Protoc Mol Biol. 2013 Jan;Chapter 22:Unit 22.1.. doi: 10.1002/0471142727.mb2201s101. PMID: 23288464; PMCID: PMC4011503.
4.News: Hundreds of drug driving convictions could be overturned due to testing error, police announce
Related Posts
How to Prevent DNA Degradation in Urine Samples?
Systematic Optimization of Urine DNA Isolation for Microbiome Sequencing
Enhance DNA Extraction from Urine Using Tris-EDTA Treatment
UTI Stops DNA Degradation in Frozen Forensic Urine Samples
Efficiency and Convenience of DNA Collection Using Buccal Swabs






